ISB-CGC BigQuery Table Search
The ISB-CGC BigQuery Table Search UI (https://bq-search.isb-cgc.org/) is a discovery tool that allows users to explore and search for ISB-CGC hosted BigQuery tables. It can be accessed directly from the ISB-CGC homepage (https://isb-cgc.org/) by clicking on Launch in the BigQuery Table Search box or selecting BigQuery Table Search from the Data Browsers drop down menu on the main menu bar.
Note: Users are not required to have a Google Cloud Platform (GCP) project or an account to learn more about the tables hosted by ISB-CGC.
Currently, ISB-CGC hosts open access BigQuery tables containing data for over 25 research programs and for over 15 data types. Each table has been curated to include detailed table and field descriptions as well as table labels allowing users to search for BigQuery tables of interest using a free-form text search or via available filters.
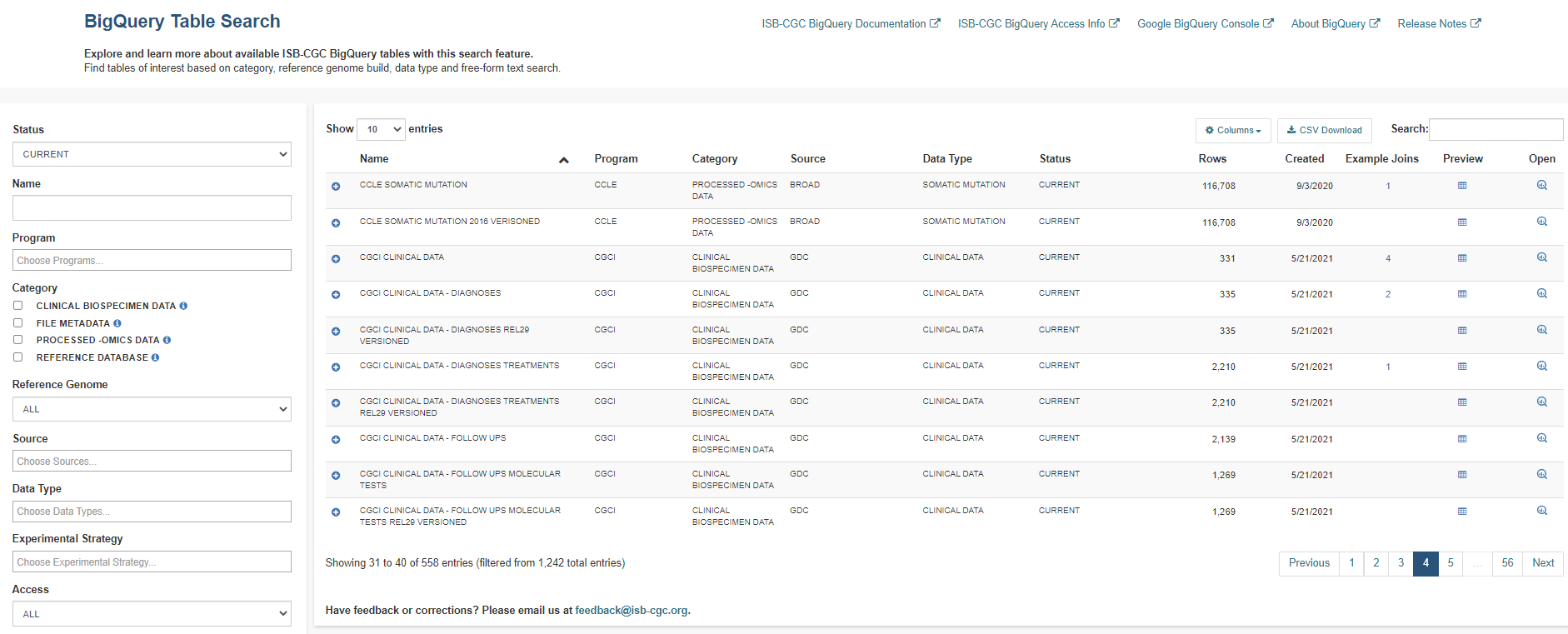
Filters
The search filters consist of a combination of multi-select dropdown lists, checkboxes and free-form text fields.
Selecting multiple items within a multi-select dropdown list or checkbox filter will perform a Boolean “AND” on those selections and bring back any data that match any of the selected items. For example, selecting Data Type BIOSPECIMEN and CLINICAL DATA will display both biospecimen and clinical data.
Selecting multiple filters will perform a Boolean “OR” on those selections and bring back only data that fits all criteria. For example, selecting Data Type BIOSPECIMEN and Source of CCLE will only display CCLE biospecimen data.
Information about each filter is detailed below.
Status
We are committed to providing the most up-to-date information in our BigQuery tables but realize that at times researchers need to reference older versions of data. Each table is assigned a status based on the following criteria:
Tables with the most up-to-date available information are given a status of current
Tables with older versions of data are given a status of archived
Tables that have data that is no longer supported are deprecated
By default, the Status filter is set to Current.
Include Always Newest tables
- When set to Off, only stable tables will display. These tables will never change. The Version is based on the version of the source data (such as GDC or PDC).
Here’s an example using filters Status: ALL, Program: BEATAML, and Category: CLINICAL BIOSPECIMEN Data and Include Always Newest tables set to Off. Only stable tables display.
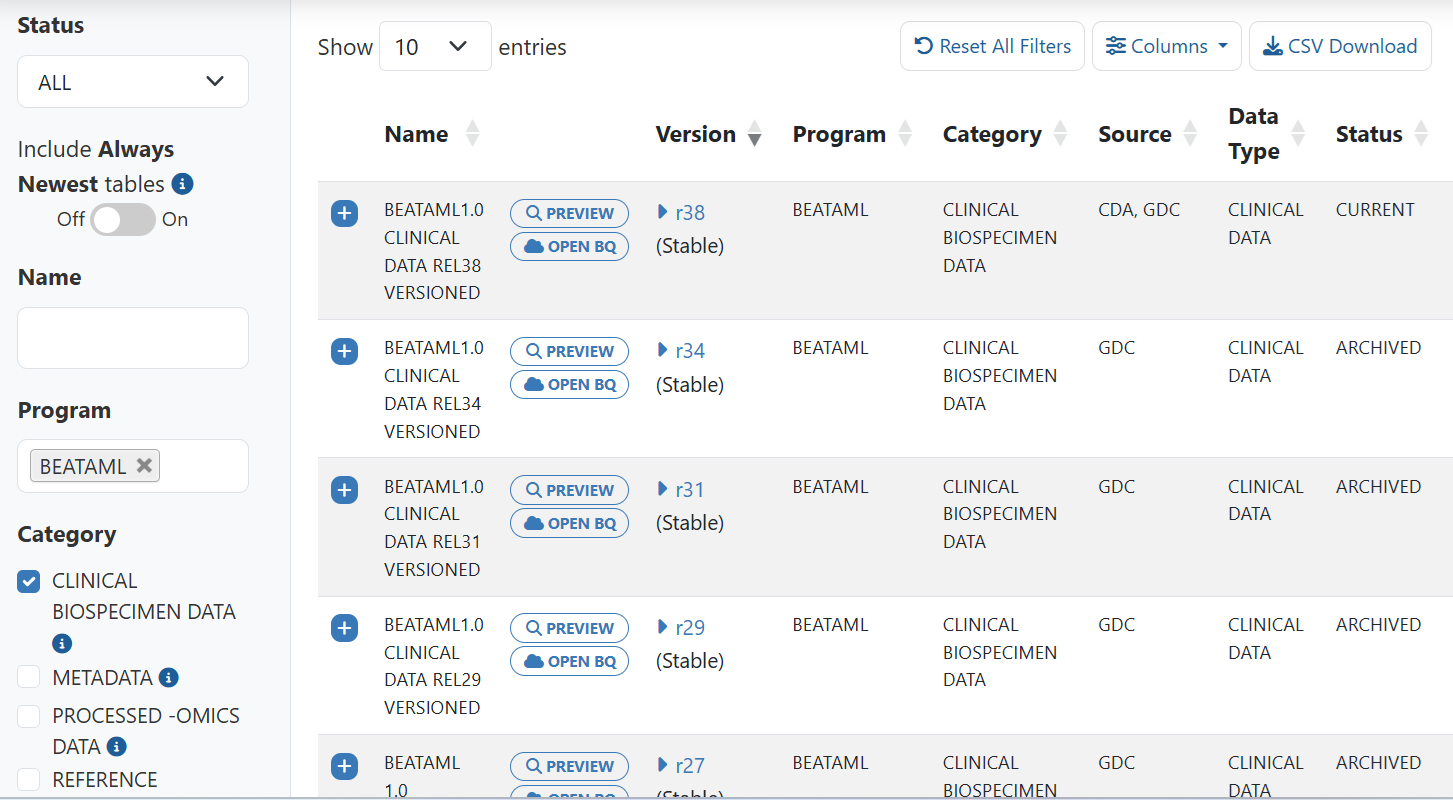
- When set to On, Always Newest tables will display along with the stable tables. These tables are a copy of the latest versioned stable table. Because they are a copy of the latest versioned table, they will be updated every time a table is created for a new version of data.
Here’s an example using filters Status: ALL, Program: BEATAML, and Category: CLINICAL BIOSPECIMEN Data and Include Always Newest tables set to On. Stable tables display but also the Always Newest table displays. In this example, this table is actually the same as the r38 versioned table. When the next version is added, it will then be the same as that table.
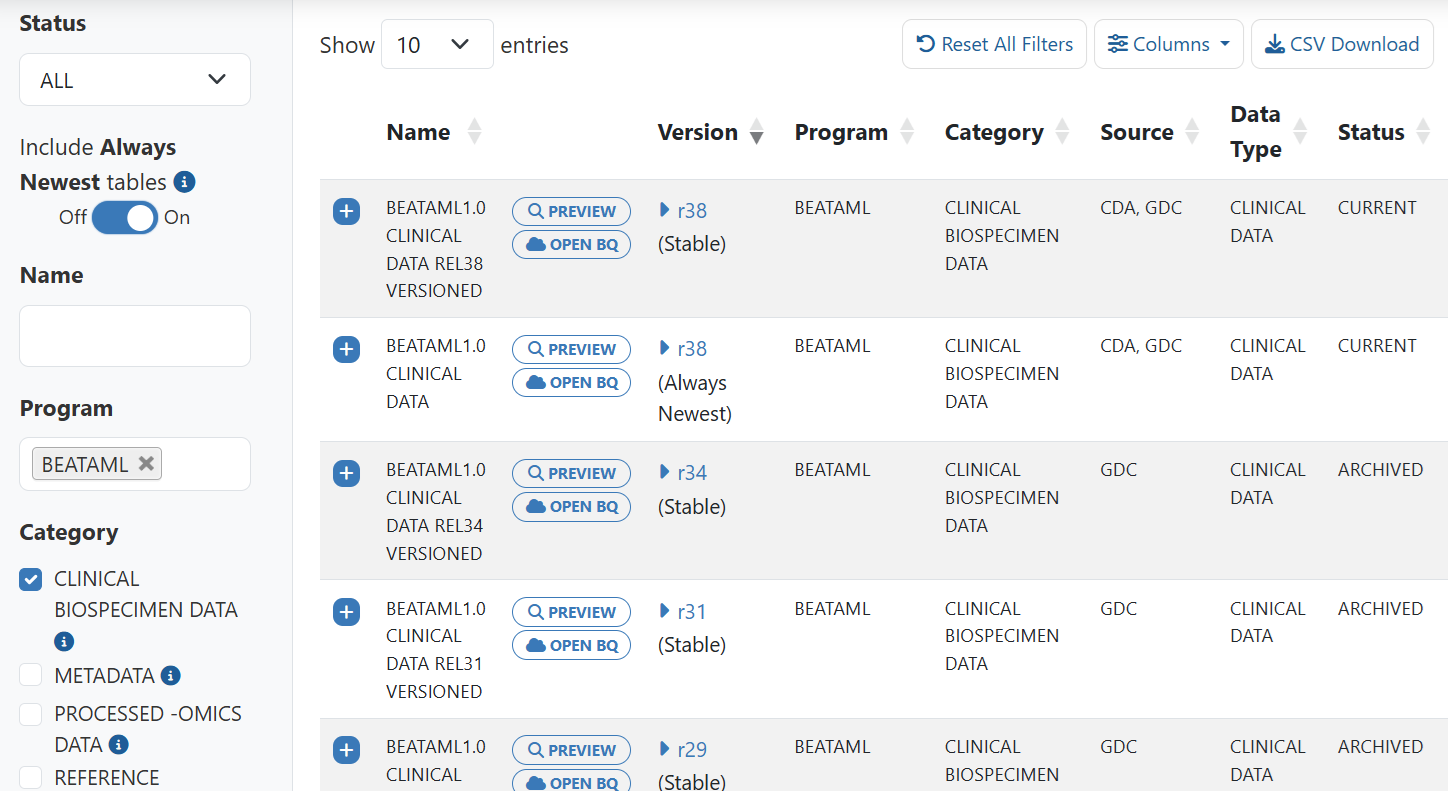
Note: Setting to On when the selected Status is ARCHIVED or DEPRECATED will not change what is displayed on the list, as archived and deprecated tables are by definition not the newest tables.
Name
The Name filter is a free-form text field; the user can type all or a portion of the name into the field to perform the search. It will match against the Name column.
Note that this Name field is not the Table ID (which is used in SQL queries) but is a Friendly Name; that is, a descriptive, user-friendly name for the table.
Program
Filter the BigQuery tables by programs such as CCLE, TARGET and TCGA by using the Program filter. Click the Program box to see the dropdown list and click on a program to select it. Additional programs can be selected by clicking in the Program box again.
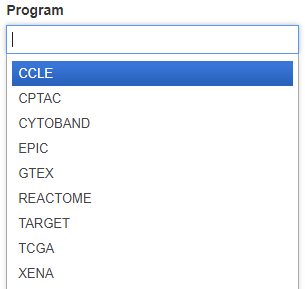
Categories
The tables are grouped into four high-level categories:
Clinical Biospecimen Data: Patient case and sample information (includes clinical tables with patient demographic data, and biospecimen data with detailed sample information)
Metadata: Information about raw data files including Google Cloud Storage Paths (includes tables with information about files available at the GDC, including GCS paths, creation dates, sizes, etc.)
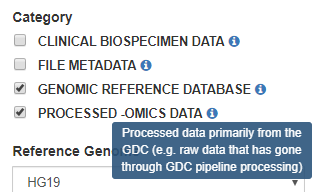
Processed -omics Data: Processed data primarily from the GDC (i.e. raw data that has gone through GDC pipeline processing e.g. gene expression, miRNA expression, copy number, somatic mutations, methylation)
Reference Database: Genomic and Proteomic information that can be used to cross-reference against processed -omics data tables (examples include ClinVar, cytoBand, dbSNP, Ensembl, Ensembl2Reactome)
Click on one or more checkboxes to select categories. Hovering the cursor over the information icon will display a short description of the category.

Reference Genome Build
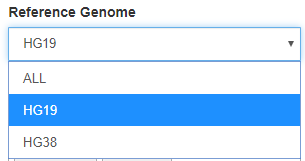
Filter for tables that contain data for hg19 or hg38. In a few cases, there are tables which contain information from both genome builds; for example, tables that include liftover coordinates between the reference builds.
By default, the Reference Genome filter is set to ALL.
Source
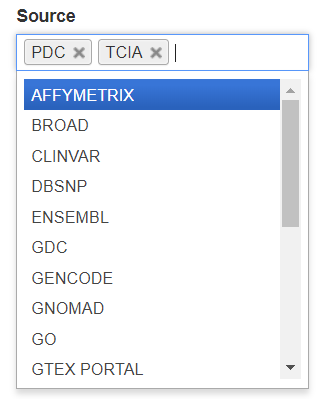
Search through the sources of the data in our BigQuery tables by using the Source filter. Click the Source box to see the dropdown list and click on a source to select it. Additional sources can be selected by clicking in the Source box again.
Data Type
The Data Type filter also allows you to filter for data types of interest. Like Source, multiple Data Types can be selected.
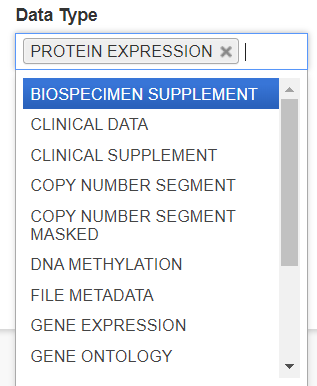
Experimental Strategy
The Experimental Strategy filter also allows you to filter for experimental strategies of interest. Multiple Experimental Strategies can be selected.
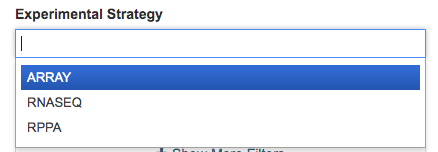
More Filters
The Show More Filters button can be used to display BQ Project, BQ Dataset, BQ Table, Table Description, Labels and Field Name filters. Except for BQ Project, these are free-form text fields; the user can type all or a portion of the name into the field to perform the query. For instance, for all datasets which have “alpha” in the name, type “alpha” into the field.
These fields are most useful for users already familiar with the BigQuery tables.
Labels
Each table was tagged with labels relating to the status, program, categoryreference genome build, source, data type, experimental strategy, access and version. Users can search on any of these labels on the Labels filter field. Users can find the Labels search filter under the Show More Filters option.
The labels for a table can be viewed when the blue plus sign (+) to the left of the table row is clicked. See the screen shot in the Schema section below.
Saving a Search Query
Each search filter and its selected value is saved as a parameter in the ISB BigQuery Table Search URL. This URL can be saved and then used again.
Here’s an example: https://bq-search.isb-cgc.org/search?status=current&category=reference_database

In this case, a Status of Current was selected, and a Category of Reference Database.
Search Results
By default, each row will display the Name, Category, Source, Data Type, Status, number of rows, and Created Date of the table.
Click on the column header to sort the displayed results by that column.
Columns Selector
Columns can be added or removed from the display by using the Columns selector. For instance, the Bq Project, BQ Dataset and BQ Table are not initially displayed, but they can be added to the display.
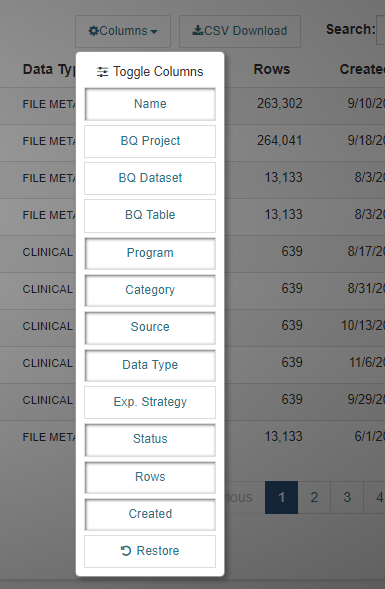
Search Box
To further filter the results, use the Search box above the results, on the right-hand side. This is a free-form text field; the user can type all or a portion of the search item into the field to perform the query. This searches all fields in the table.
Export
To export the results of your search to a file in Comma Separated Values (CSV) format, click the CSV Download button.
Schema Description
For detailed table information, click on the blue plus sign (+) on the left-hand side.
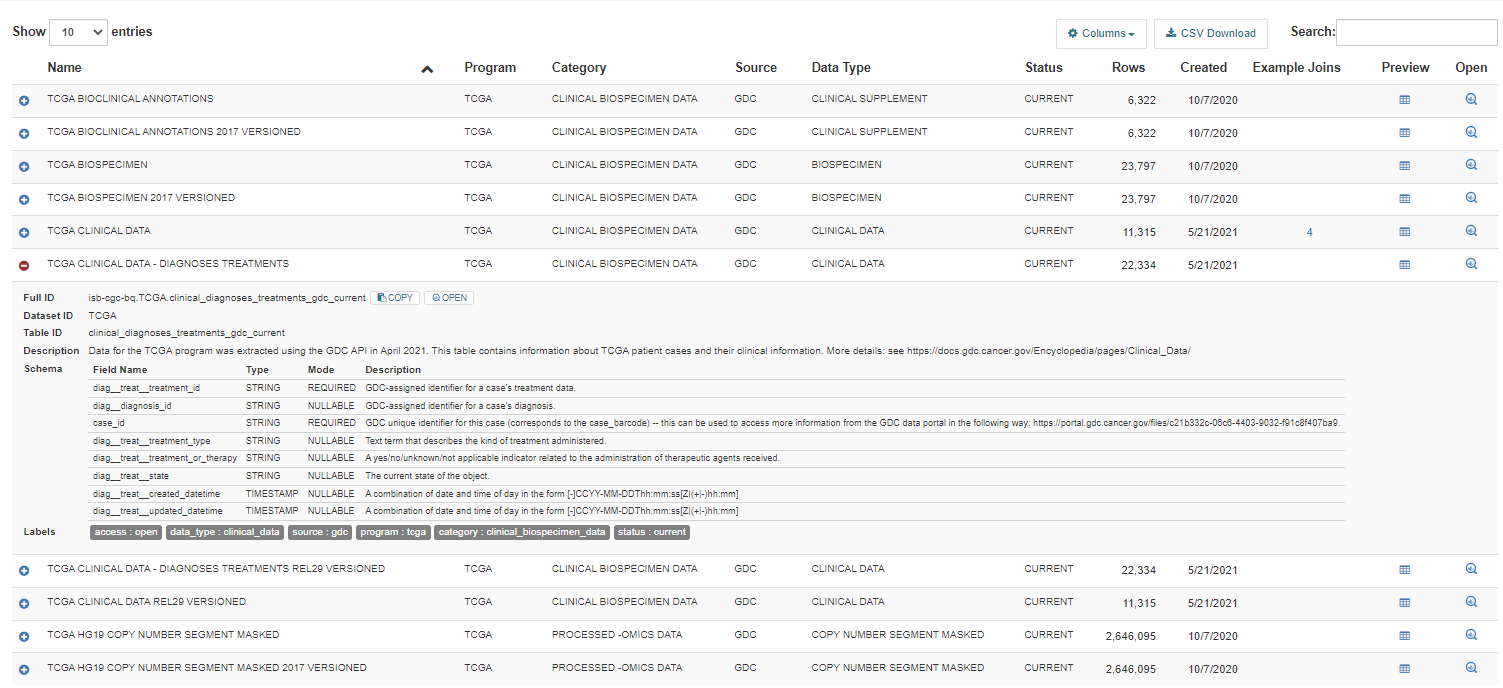
The following information is displayed:
Full ID - This is the Project, Dataset ID, and Table ID concatenated with periods between them. The Full ID is used in SQL queries.
Dataset ID - The BigQuery dataset of the table. A data set is a group of related tables.
Table ID - The BigQuery table ID.
Description - A description of the table, which includes information such as how the data was created, its source, data type, and contents.
Schema - The schema displays the Field Name, Type, Mode and Field Description for each field in the table.
Labels - Labels are table metadata describing the source, category, program, data type, reference genome build, status, version and access of the table data.
Copy button
Next to the Full ID is a Copy button. When the user clicks this, the Full ID is copied to the clipboard. The Full ID can then be pasted into an SQL query within the BigQuery Query editor.
Open button
Next to the Copy button is an Open button. Clicking on this button opens the table in the BigQuery Google Cloud Platform Console. For more details, see the Table Access in Google BigQuery section below.
Table Preview
A few rows of the data in a BigQuery table can be viewed by clicking on the Preview button. This feature allows the user to get a better idea of the contents and format of the data.
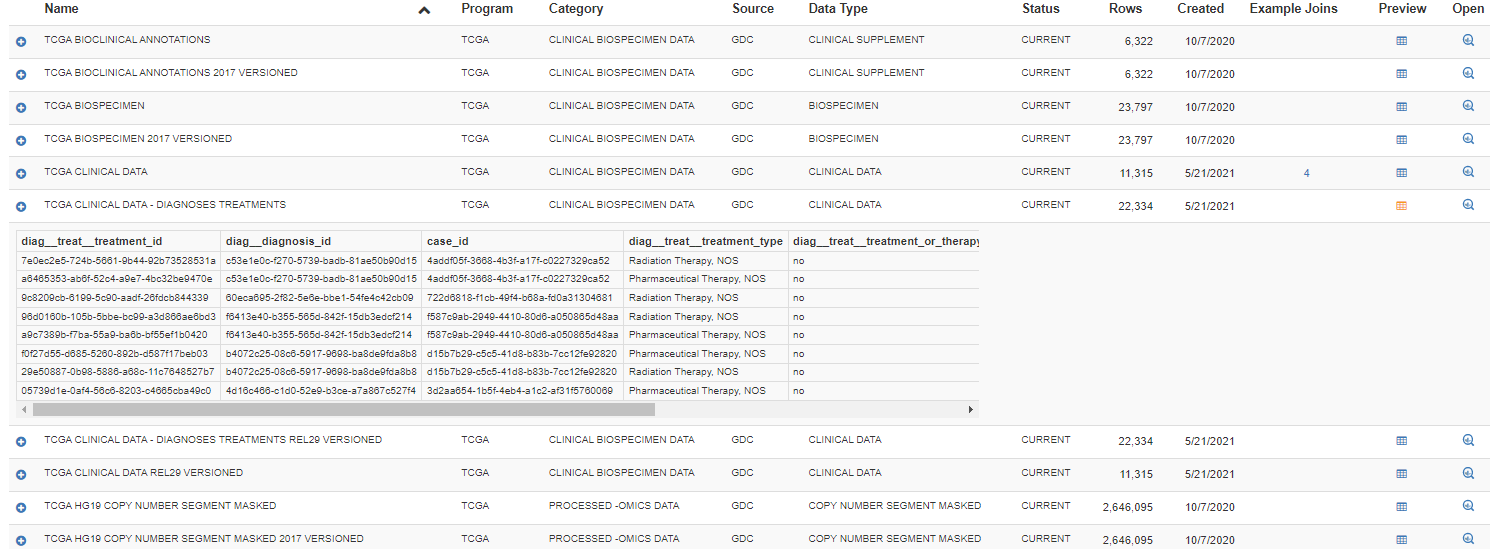
Example Joins
The Example Joins column specifies the number of example SQL join queries, for the table on that row, which are provided by the BigQuery Table Search. Clicking on the number will display a list of the examples.
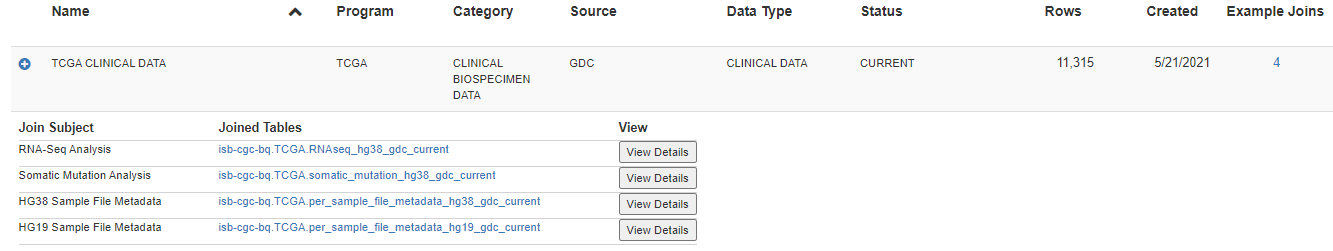
The following information is displayed:
Join Subject - This is the topic of the query.
Joined Tables - Here, the tables which are joined with the table in the row are listed.
View - The View Details button takes you to a screen which displays the SQL statement and a more detailed description of the query.
Join Details
Clicking on the View button displays the Join Details screen.
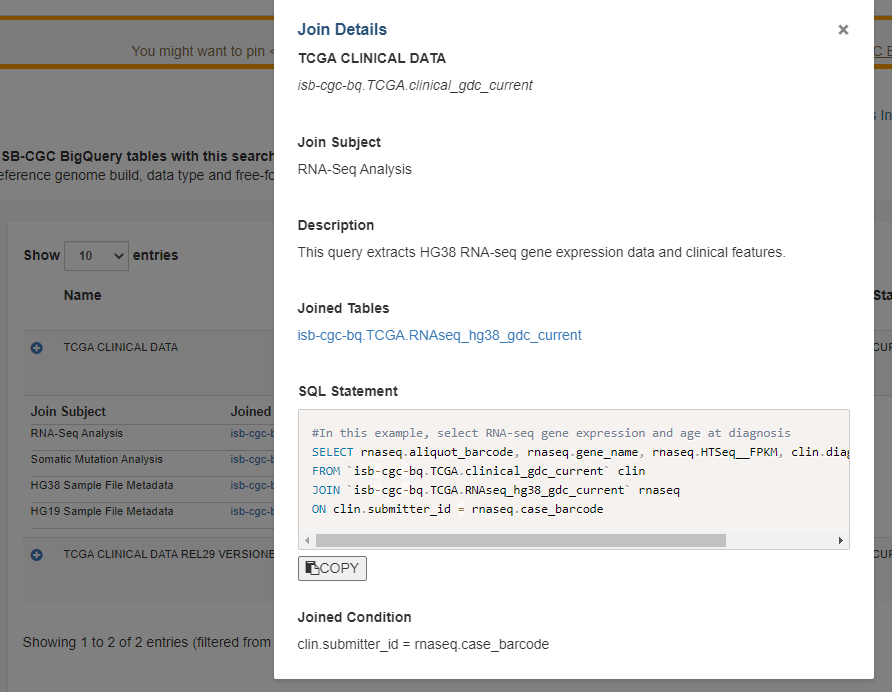
The following information is displayed:
Table identification - Both the table name and the table Full ID are displayed.
Join Subject - This is the topic of the query.
Description - Here, the query is described in more detail. For instance, it will describe what kind of data is extracted.
Joined Tables - Tables which are joined with the main table of interest are listed here. The table name is also a link, in case you would like to easily learn more about the joined table. Clicking on it will open up the ISB-CGC BigQuery Search in another tab, with the table information in the query results.
SQL Statement - This is the SQL statement for the joined tables.
COPY - Clicking this button copies the SQL Statement to your clipboard. You can then directly copy the SQL query into the Google Cloud Platform BigQuery Console, a Jupyter notebook, or anywhere that you would like. These queries can be run as they are, or you can tailor them to your needs.
Joined Condition - There are the fields being joined between the tables.
Table Access in Google BigQuery
To access the BigQuery tables in Google Cloud Console directly from the Table Search UI, click on the Open BQ button.
Note:
If you have previously accessed the Google Cloud Platform and have a Google Cloud Platform project already set up, this button will automatically open up the table in the Google BigQuery Console as depicted in the image below.
If you have never accessed Google Cloud Platform, you will be presented with a Google login page. You can use any Google ID to log in. Instructions on how to create a Google identity if you don’t already have one can be found here. You will be prompted to create a project, free of charge. Once you create the project, you will be directed to the BigQuery table you wished to open in the Google BigQuery Cloud Platform Console.
Google Cloud Platform’s free tier allows users to access many common Google Cloud resources including BigQuery free of charge and query up to 1 TB of data per month for free.
Please see the following ISB-CGC documentation pages for guidance:
